Web demos
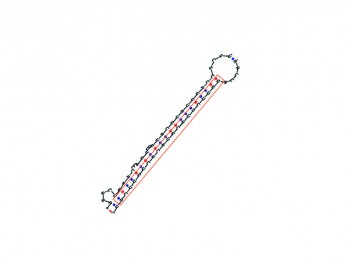
miRe2e
An end-to-end deep learning model based on Transformers for prediction of pre-miRNAs.
DL4papers
A deep learning approach for the automatic interpretation of scientific articles.
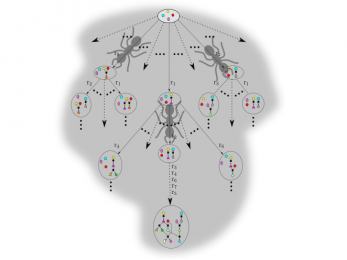
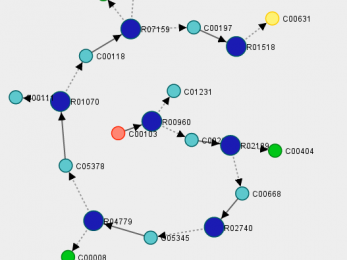
PhDSeeker
PhDSeeker is a bio-inspired algorithm for synthesizing linear and branched metabolic pathways.
Beta Dereverberation
A blind, single channel dereverberation method in the time-frequency domain.
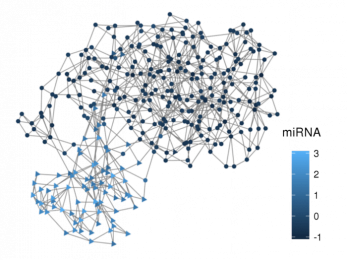
miRNAss
Semi-supervised method for microRNA prediction with few labelled samples.
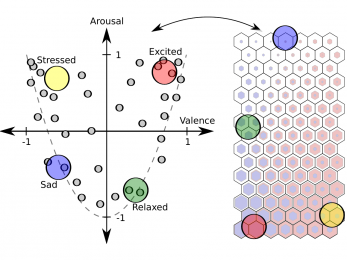
Dimensional affect recognition
Arousal and valence are estimated from features of heart rate variability.
miRNA: supervised vs unsupervised
Compares a supervised vs an unsupervised approach for pre-miRNA prediction in model genomes.
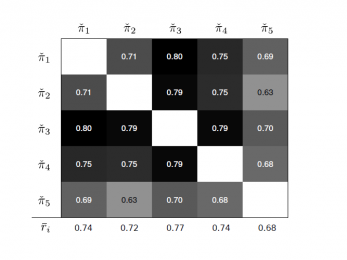
divControl
A novel method to smoothly control the diversity of a cluster ensemble.
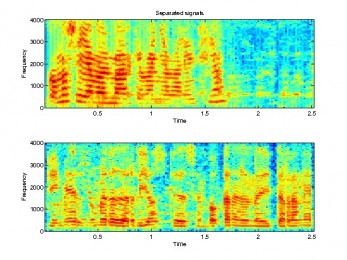
multibinICA
Algorithm for frequency-domain blind source separation based on Multibin ICA for 2 by 2 mixtures.
fdICA
Algorithm for frequency-domain blind source separation based on the pseudoanechoic model.
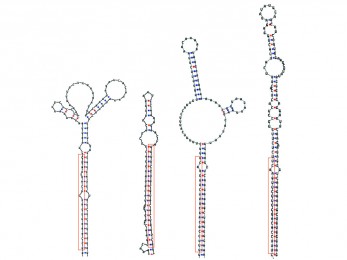
miRNAfe full
miRNAfe full is an advanced tool to extract features from RNA sequences, providing almost all state-of-the-art feature extraction methods published today.
bSOM lite
Simple demo of bSOM, an algorithm for improve clustering with information from metabolic pathways.
NRFAR
An algorithm designed to recognize foraging behavior of cattle in free-grazing environments
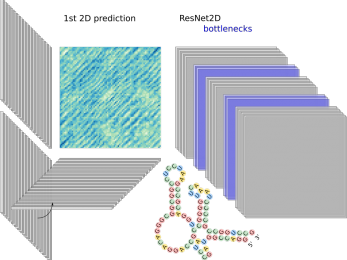
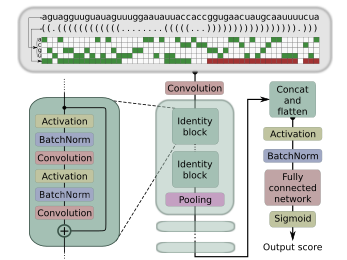
mirDNN
A convolutional deep residual neural network for prediction of pre-miRNAs in genome-wide data.
BUFAR
An online algorithm to extract detailed information of foraging behavior in grazing cattle.
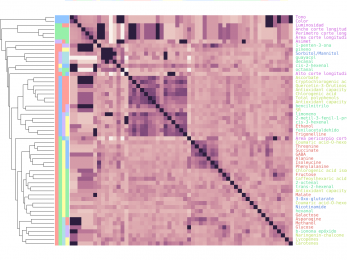
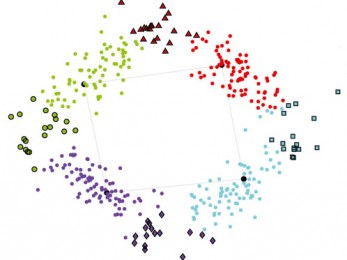
Clustermatch
Clustermatch is an efficient clustering method able to process highly diverse datasest.
RAFAR
A regularity-based algorithm designed for long-term analysis of foraging behavior in grazing cattle.
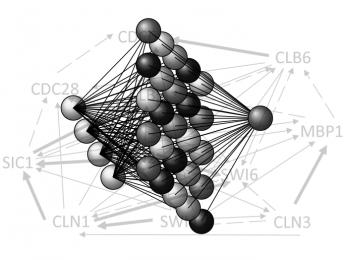
ELM-GRNNminer
Extreme learning machines for reverse engineering of gene regulatory networks from expression time series.
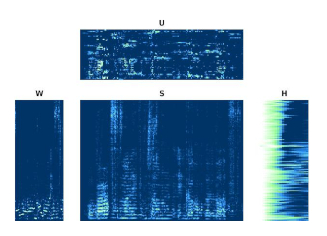
blindder
Blind speech dereverberation based on convolutive non-negative matrix factorization with mixed penalization.
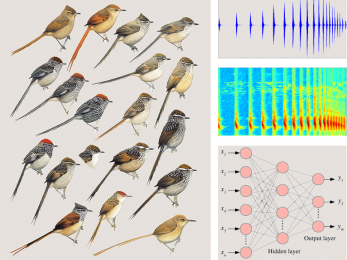
Furnariidae recognition
Classification of Furnariidae species using speech-related features and machine learning.
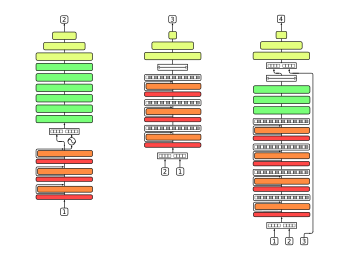
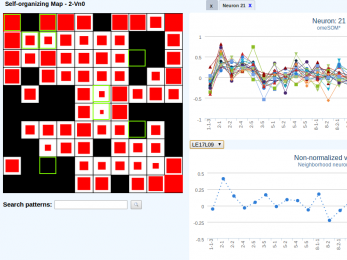
deepSOM
A novel and effective way of approaching high class-imbalance in pre-miRNA prediction.
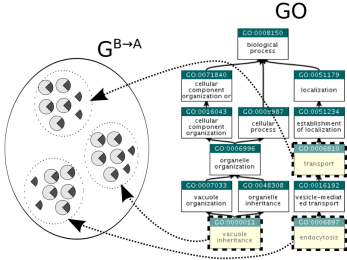
gamma-AM
A novel method for inferring biological function for a set of genes with previously unknown function.
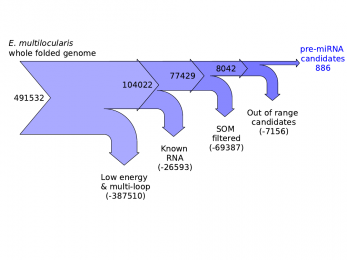
miRNA-SOM
miRNA-SOM is a tool for the discovery of pre-miRNA in the E. multilocularis genome.
EvoMS
An evolutionary tool for finding novel metabolic pathways linking compounds through feasible reactions.
miRNAfe
A comprehensive tool providing almost all state-of-the-art RNA feature extraction methods used today.
omeSOM full
*omeSOM is a tool designed to give support to the data mining task of metabolic and transcriptional datasets derived from different databases.